Our Diagnostics Portfolio
What is Panel Diagnostics?
Our panel diagnostics assists you to secure a clinical diagnosis, make a prognostic assessment of the disease, and allow early therapeutic intervention. Furthermore, based on the findings of the index patient, targeted investigation of family members can be offered.
To maximize diagnostic yield, all diagnostic panels include a CNV analysis and are based on our unique CeGaT ExomeXtra®, which enriches not only coding but also all diagnostically-relevant intronic and intergenic variants.
Key features of our panel diagnostics:
- sequencing of ExomeXtra®
- analysis of single nucleotide variants (SNVs), small insertions and deletions (indels) and copy number variants (CNVs)
- known pathogenic intronic variants and phenotype associated mtDNA hotspots included
- full mtDNA analysis possible
- chance of identification of mosaicisms
- high average sequencing coverage: > 100x (Illumina Platform)
- medical report written by a team of experts incl. medical doctors specialized in human genetics
- turnaround time: 4 – 6 weeks
(Due to high demand and a large number of urgent and prenatal cases, our turnaround time is currently slightly increased. For urgent samples, our processing time will of course remain 2-3 weeks.)
CeGaT’s ExomExtra® Sequencing and Large Panel Approach
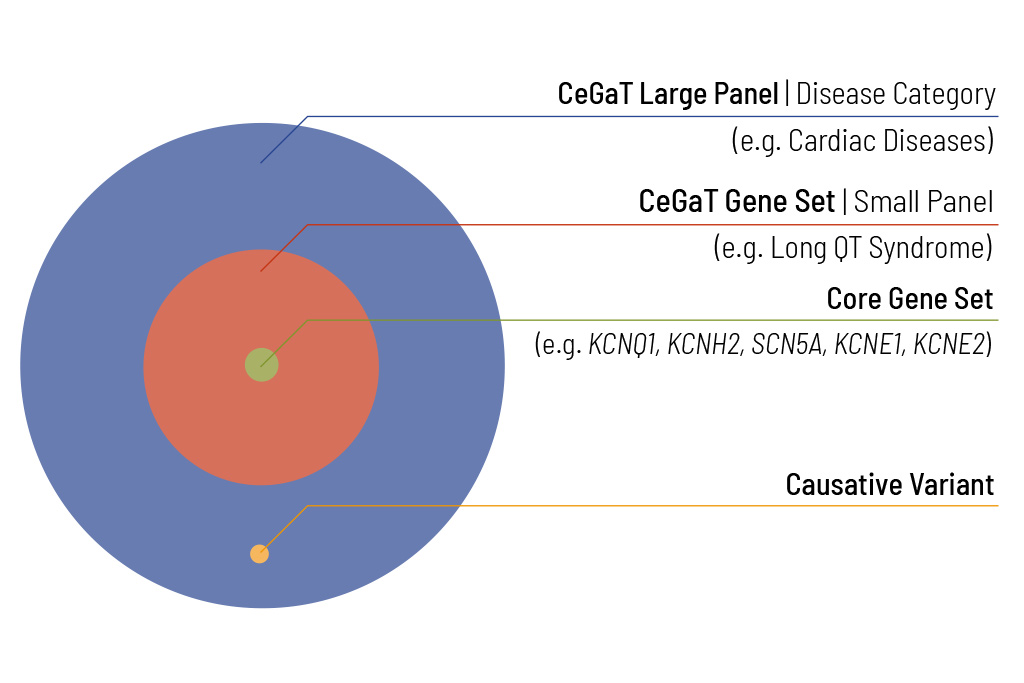
Identifying the genetic cause of the patient’s disease is the ultimate goal of our diagnostic panels. For many diseases, the number of potentially disease-causing genes is very large. To enable efficient diagnostics for these diseases, our diagnostic panels are evaluated based on ExomeXtra®. Sequencing with ExomeXtra® covers all clinically relevant regions throughout the genome and considers factors that are often overlooked in regular genetic testing. By covering all known pathogenic intronic and intergenic variants in addition to all protein-coding regions, CeGaT’s ExomeXtra® provides an unmatched basis for the best genetic diagnostics.
We offer to expand the analysis to the complete diagnostic panel for pathogenic (ACMG class 5) and likely pathogenic (ACMG class 4) variants of the large panel relevant to differential diagnoses.
CNV Analysis
Similar to single nucleotide variants (SNVs) and small insertions and deletions (indels), copy number variants (CNVs) can cause the patient’s phenotype. Hence genetic testing without CNV analysis is incomplete and may lead to a negative medical report even though there is a genetic cause. At CeGaT a CNV analysis is an integral part of our Diagnostic Panels; it does not need to be requested, it is included in every Diagnostic Panel.
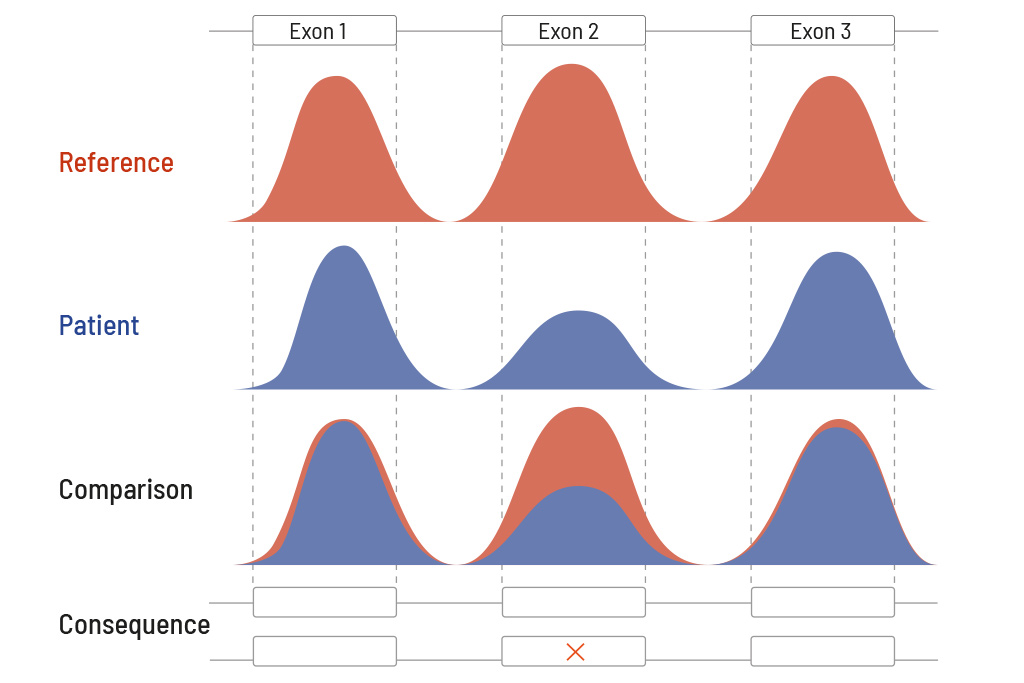
Technically speaking, CeGaT’s CNV analysis is based on a reference map for the expected coverage. This reference map is created from our large in-house database of diagnostic cases. The database comprises a high five-digit number of diagnostic cases. The individual patient sample is compared with this reference map (see figure on the right). Due to the high number of cases in our database, the sensitivity of the CNV analysis is >81% for single exon deletions and >96% for three or more exon deletions. Identified and reported CNVs are validated by MLPA or qPCR.
To ensure best possible diagnostic yield, all our Diagnostic Panels include the analysis of single nucleotide variants (SNVs), small insertions and deletions (indels), copy number variants (CNVs), known pathogenic intronic variants and phenotype associated mtDNA hotspots!
Our Accreditations
Binding standards guarantee the quality of our work: Our laboratory services are accredited according to CAP/CLIA and DIN EN ISO 15189. You can find further accreditations and certifications here.
You Are also Welcome to Take a Look at the Following Areas
Contact Us
Do you have a question, or are you interested in our service?
Diagnostic Support
We will assist you in selecting the diagnostic strategy – for each patient.